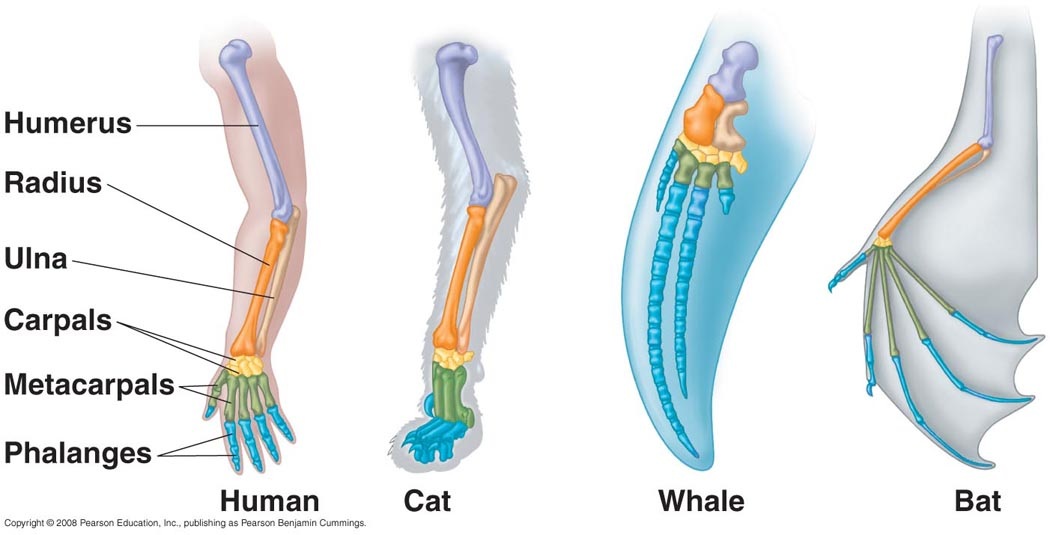
What is homology?
Homologous proteins are derived from a common amino acid sequence, but may have developed new functions. These proteins might have small sequence changes that alter amino acid structure, but overall one would likely expect higher percent identity in more related species and less in more distant. If a protein maintains the same function within both organisms, the protein is orthologous. Some proteins are highly conserved throughout evolution. These proteins often serve important cellular maintenance or structural functions.
Protein homology can also be important when selecting an appropriate model organism for research. If your protein is highly conserved and reveals a similar phenotype across species, many model organisms might suit your research. If your protein is less conserved, percent identity and conservation of domains may play an important role in choosing an appropriate candidate. |
|
Alignment Tools: BLAST and Homolgene
To identify homologous proteins between organisms, tools like NCBI's BLAST (Basic Local Alignment Search Tool) compares input sequences (DNA, mRNA, or protein) to a database of known sequences and genomes and provides top matches based on how identical the sequences are [1]. NCBI's Homologene similarly creates a list of top matches for a known protein, but returns only one per organism [2]. Additionally, some data on comparative domain structure is available through the Homologene assessment. Pyrin Homology Analysis
As predicted from the MaxID of sequences and known evolutionary relationships between species, Homo sapiens pyrin isoform 1's top matches in Homologene were other primate species, followed by other mammalian species (see Protein References below). Other model organism species had relevant homologous proteins, but were much less conserved than previous matches. All vertebrate homologous proteins shared a tri-partite domain protein (TRIM) structure (Domain Page), and shared many of the same functional domains.
Canis lupus familiaris was surprisingly well-conserved in sequence homology and appears to conserve domain structures, and could prove to be a valuable model organism for pyrin. Mouse models have been shown to have the same phenotype, despite lacking a functional SPRY domain (Model Organisms). |
Figure 2. Homologene alignment for Homo sapiens pyrin isoform 1.
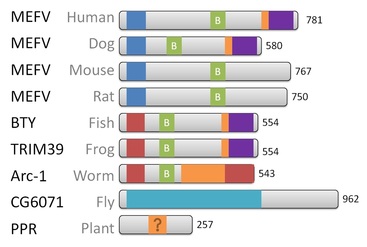
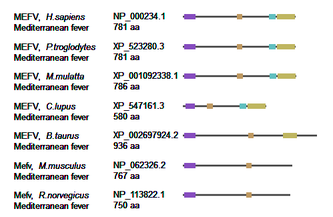
Figure 3. Protein homology visualization of MEFV in common model organisms.
|
Homologous Protein ReferencesHomologous proteins were identified through a BLAST search and compared to Homologene results [1,2]. Specific model organisms were searched for by organism and the highest Max ID match was chosen as a homologous protein. All proteins were BLAST-ed against Homo sapiens pyrin isoform 1 on "Highly similar sequences (megablast)."
|
|
Homologene Matches
Homo sapiens - pyrin isoform 1 Accession Number: NP_000234.1 FASTA Pan troglodytes - pyrin isoform 2 Accession Number: XP_523280 E-value: 0.0 Max ID: 98% FASTA Macaca mulatta - pyrin Accession Number: XP_001092338.1 E-value: 0.0 Max ID: 88% FASTA Canis lupus familiaris - pyrin Accession Number: XP_547161.3 E-value: 0.0 Max ID: 75% FASTA Bos taurus - pyrin Accession Number: XP_002697924.2 E-value: 0.0 Max ID: 51% FASTA Mus musculus - Mediterranean fever isoform CRA_b Accession Number: EDK97204.1 E-value: 2e-173 Max ID: 52% FASTA Rattus norvegicus - Mediterranean fever isoform CRA_b Accession Number: EDL96327.1 E-value: 2e-141 Max ID: 46% FASTA |
Other Common Model Organisms
Danio rerio - Bloodthirsty Accession Number: NP_001018311 E-value: 3e-57 Max ID: 34% FASTA Xenopus (Silurana) tropicalis - TRIM 39 Accession Number: NP_001123789.1 E-value: 4e-59 Max ID: 34% FASTA Drosophila melanogaster - CG6071 Accession Number: NP_648488.1 E-value: 0.050 Max ID: 24% FASTA Caenorhabditis elegans - Protein ARC-1 Accession Number: CAA87044.3 E-value: 0.060 Max ID: 29% FASTA Arabidopsis thaliana - PPR protein Accession Number: NP_175000.1 E-value: 2.6 Max ID: 28% FASTA |