Protein Interaction Networks
Creating Protein Interaction Networks
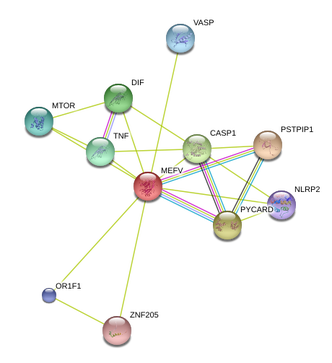
Proteins rarely act in isolation, but instead complex and act in signal transduction pathways to accomplish their biochemical function. Large-scale databases of protein-protein interactions can provide useful information about pleiotropic effects, effectors in drug toxicity, and provide hints about biological function for uncharacterized proteins [1]. To investigate protein-protein interactions on the global scale, techniques such as two-hybrid systems, TAP tag, protein microarrays, mass spectrometry and CHiP assays have all been used [1]. The result: many huge databases able to churn out protein interactions like the ones shown in Figure 1 and 2. One available database of known and predicted protein interactions is the STRING database. This database derives both direct, physical interactions and indirect, function associations from genomic context, high-throughput experiments, coexpression, and primary literature. STRING will be used in the following analysis. Human Pyrin Pyrin has a number of known interactors associated with caspase-mediated interleukin production (CASP1, PYCARD, NLRP2), stimulation of interleukins (TNF, DIF), and actin-associated proteins (VSP, PSTPIP1). It also interacts with MTOR (a kinase subunit which regulates cell growth and surivival), ORF1 (an olfactory receptor) and ZNF205 (a zinc-finger protein). Comparison with Zebrafish btr26 A well-conserved protein-protein interaction can provide interesting information about the evolution of one's protein through different species. Though pyrin function is well-conserved throughout mammals, any protein species preserved through zebrafish would identify an interesting target gene. PSTPIP1 is the only gene preserved that interacts with the homologous zebrafish bloodthirsty (btr26) gene. |
Figure 1. Human MEFV and it's protein functional associations and binding interactions. From STRING.
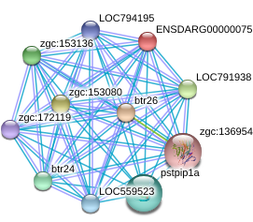
Figure 2. Zebrafish btr26 and it's protein functional associations and binding interactions. From STRING.
|
|
Analysis
Protein-protein interactions solidify the information already known about pyrin's biochemical processes in the cell. Additionally, protein-protein interaction databases provide a wealth of information about potential effectors and pathways that may be affected from mutations or new drug treatments. PSTPIP1 provides an interesting protein interaction that should be further examined for its role in response to pyrin. References [1] Pellegrini, M, Haynor, D, and Johnson, JM. (2004) Protein interaction networks. Expert Rev. Proteomics 1(2):89-99. |