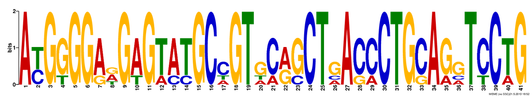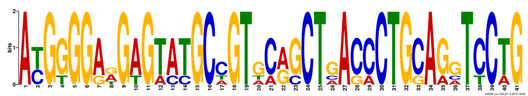What are DNA motifs and domains?
A common misconception is that DNA motifs and domains are referring to the same structures and patterns. A DNA motif is a short, recurring pattern in DNA that is assumed to have a biological function. These could be indicative of common functional binding domains, and therefore indicative of a similar function. Protein domains are sequences subject to pressures to maintain functional interactions. These may often result in specific interactions with DNA, other proteins, small molecules and metal ions.
|
|
DNA Motifs
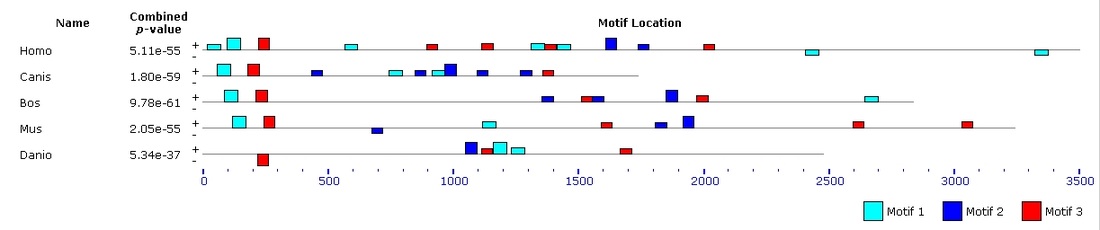
There are several methods for identifying DNA motifs and protein domains. These include enumeration, deterministic optimization, and probabilistic optimization. Enumeration is a very broad method which looks at all possible motifs in the entire search space, however they are limited in defining more subtle sequence patterns [1]. A deterministic optimization model uses an expectation maximization algorithm, which takes small segments of sequence and compares the probability that it was generated by the motif relative to background sequence [1]. MEME is a program that uses this approach. Lastly, a probabilistic optimization uses Gibbs sampling approach and uses randomized implementation of the expectation maximization algorithm [1]. MEME online tools were used to compare the human variant 1, dog, bull, mouse and zebrafish homologs and search for DNA motifs [2]. The results are shown in Figure 1, 2, and 3. Figure 4 depicts the alignment between MEFV homologs across the region identified as Motif 1. The first four organisms have near-identical matches in these regions, whereas zebrafish has the most basepair changes. Motif block diagrams show the order and spacing of non-overlapping matches to the motifs in each high-scoring sequence. Each motif is labeled with the full square of each color being the best match for each motif in the sequence. Because of the variation in species-specific length of the sequence and intron location, not all motifs were aligned when DNA sequences are compared next to each other. |
Figure 1. Motif 1, E-value of 2.8e-027
Figure 2. Motif 2, E-value of 1.1e-016
Figure 3. Motif 3, E-value of 1.7e-015
Figure 4. Alignment of MEFV homologs in Homo, Canis, Bos, Mus and Danio for segment of Motif 1.
Figure 5. Motif block diagram of MEFV homologs in Homo, Canis, Bos, Mus and Danio.
|
|
Analysis
The results above, obtained from MEME, show three different DNA motifs present in the MEFV gene. These results are simple to interpret, where the height of each nucleotide reveals how common it was between the sequences analyzed. Additionally, MEME allows for multiple nucleotides to be listed in any single slot. Where there is only one large nucletoide, that one was present in all of the sequences, whereas a location where multiple short letters are listed was not as well conserved. All of the motifs found when comparing human variant 1, dog, bull, mouse and zebrafish were 41 or 50 basepairs long, which makes them highly viable results. In Figure 4, the alignment of a single motif between species is identified. One can see that the Canis, Bos, Homo and Mus all align very well with only a few amino acid substitutions. If Danio had been excluded from this analysis, the motifs would likely have been better conserved. |
References
[1] D'haeseleer, P. (2006) How does DNA sequence motif discovery work? Nature Biotechnology. 24(8), 959. doi: 10.1038/nbt0806-959
[2] Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
[1] D'haeseleer, P. (2006) How does DNA sequence motif discovery work? Nature Biotechnology. 24(8), 959. doi: 10.1038/nbt0806-959
[2] Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.